Last Updated:
Process Illumina methylation data in BRB-ArrayTools
BRB-Arraytools imports Illumina Methylation data using the R package "lumi". Beta values, reflecting the proportion of methlyated probes and ranging from 0 to 1, will then be normalized and transformed to M values, which are similar to log2(beta/(1-beta)) values. Different analysis tools, such as class comparison, prediction, hierarchical clustering, and enrichment analysis, can be conducted using the transformed and normalized M values. In addition to the tools shared with other types of microarray data, analysis tools have been developed to find frequently methylated regions and to correlate methylation with expression data in BRB ArrayTools.
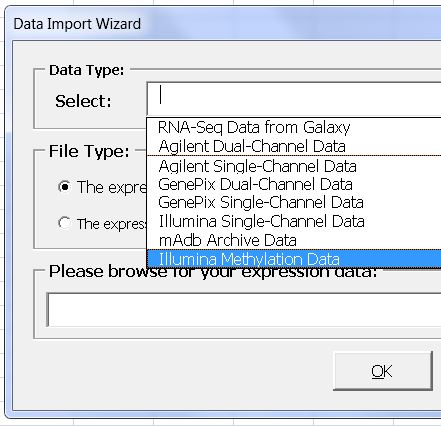
Find frequently methylated region in BRB-ArrayTools
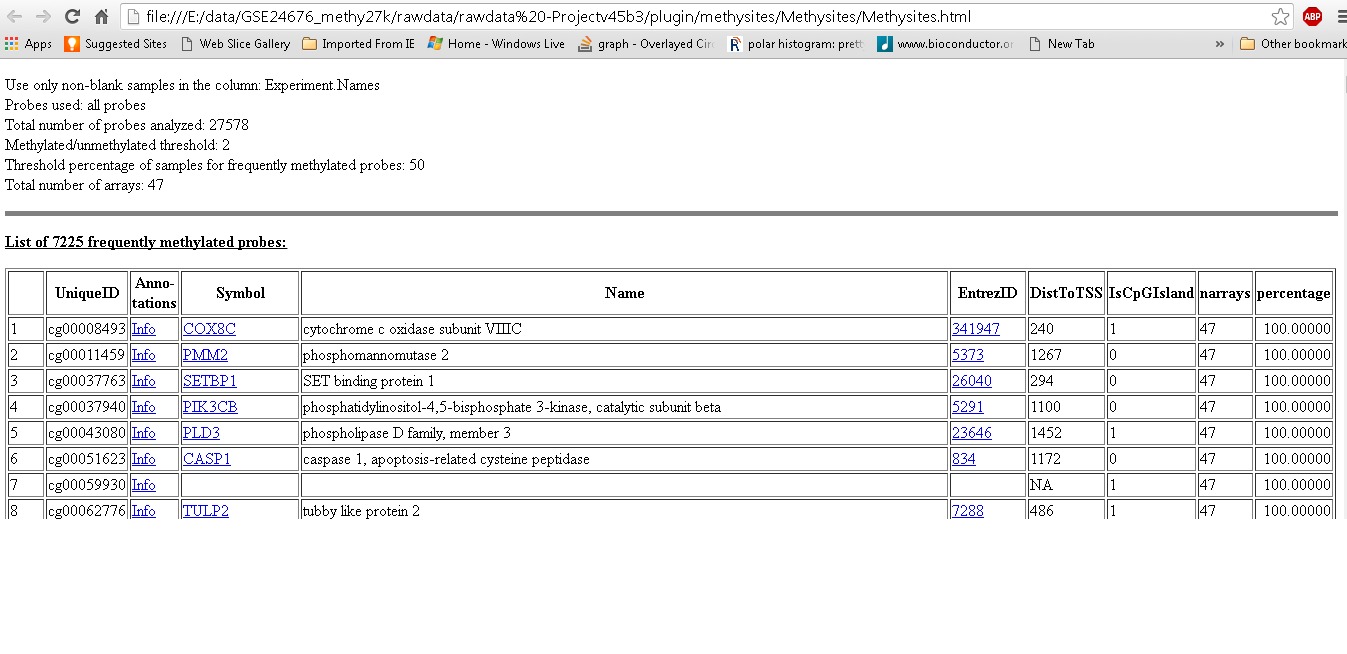
Create a heat map with methylation data